The evaluation of Bcftools mpileup and GATK HaplotypeCaller for variant calling in non-human species | Scientific Reports
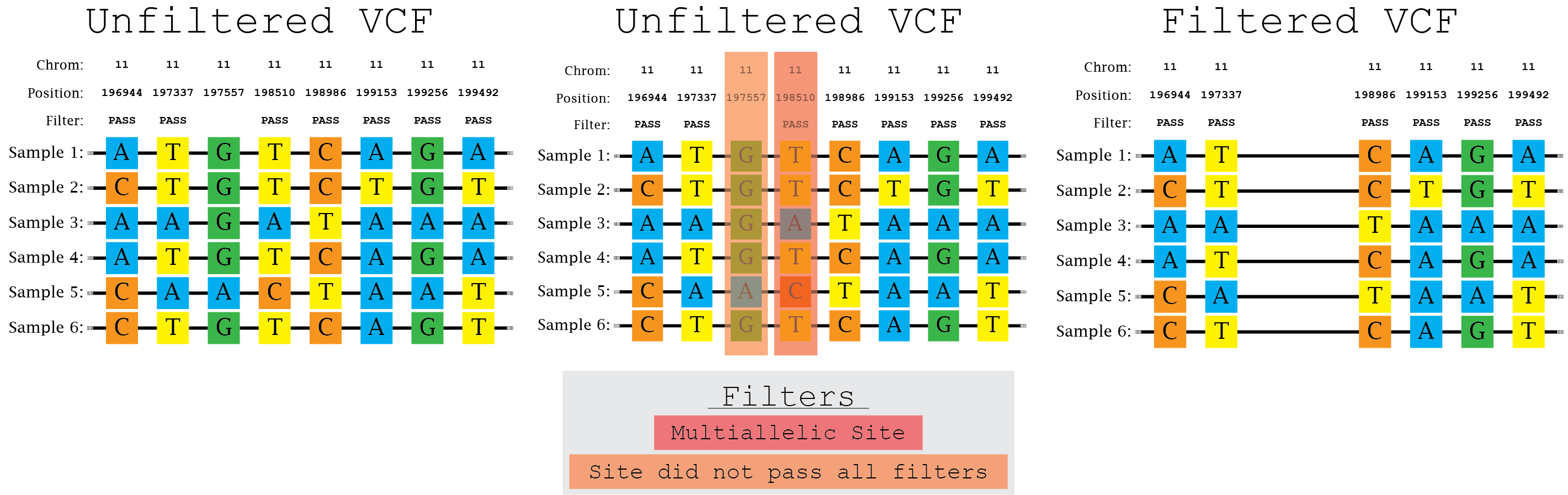
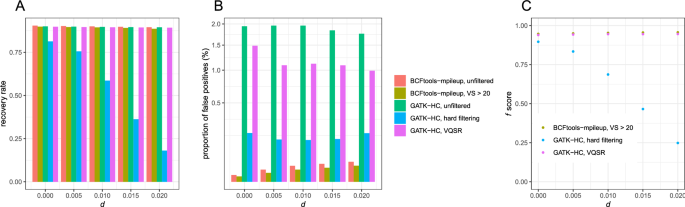
a) Filtering different variant callers VCF output for SARS-CoV-2 data.... | Download Scientific Diagram
The evaluation of Bcftools mpileup and GATK HaplotypeCaller for variant calling in non-human species | Scientific Reports

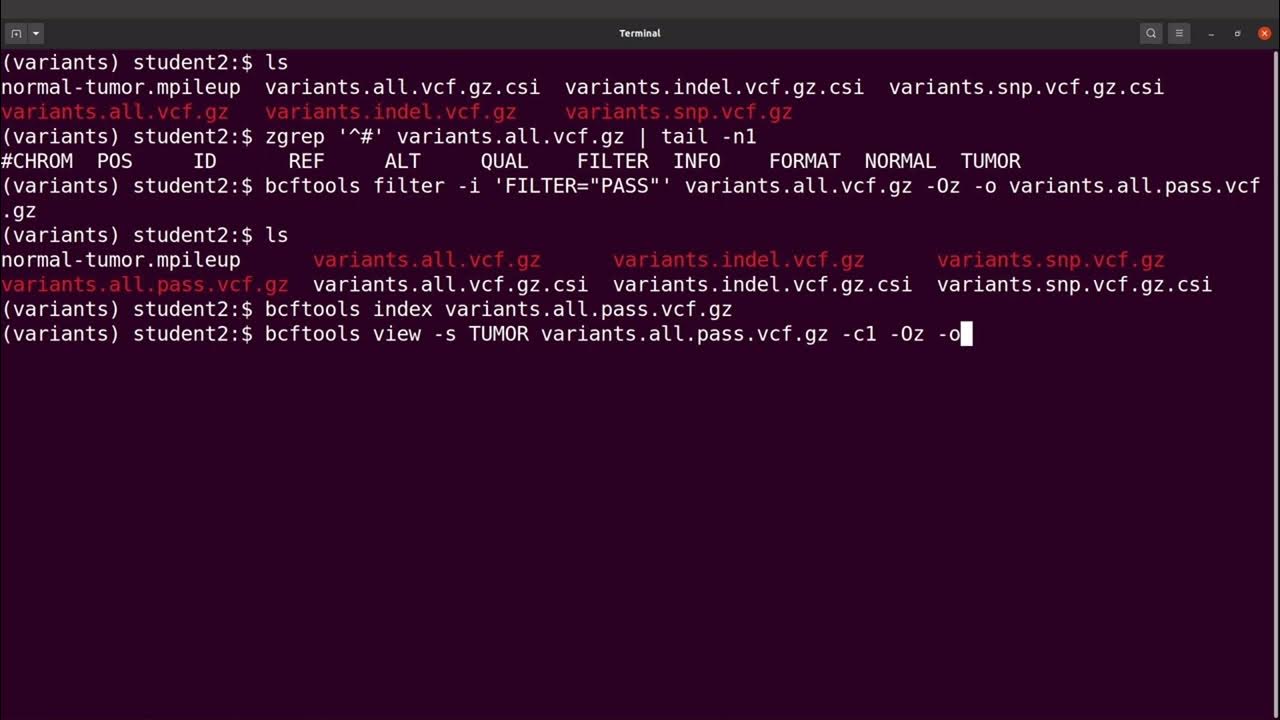
bcftools filter -e 'GT="het"' recognize GT 2/2 as heterozygous · Issue #1268 · samtools/bcftools · GitHub

PDF) How to extract and filter SNP data from the genotyping-by- sequencing (GBS) data in vcf format using bcftools