Genes | Free Full-Text | A Novel Function for the Conserved Glutamate Residue in the Walker B Motif of Replication Factor C

The Conserved Glutamate Residue Adjacent to the Walker-B Motif Is the Catalytic Base for ATP Hydrolysis in the ATP-binding Cassette Transporter BmrA* - Journal of Biological Chemistry

Walker-A Motif Acts to Coordinate ATP Hydrolysis with Motor Output in Viral DNA Packaging - ScienceDirect
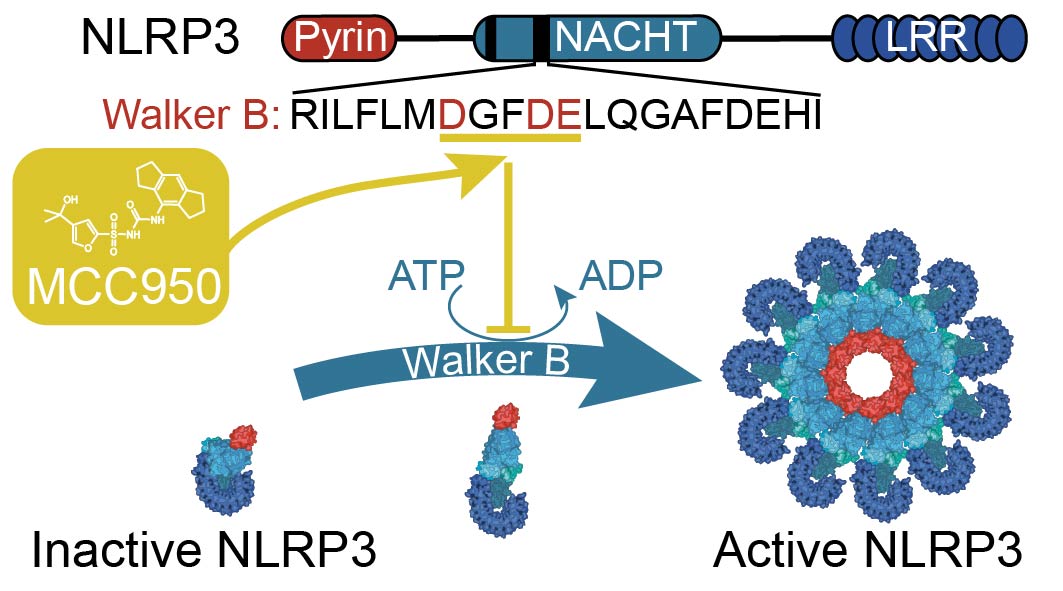
MCC950 directly targets the NLRP3 ATP-hydrolysis motif for inflammasome inhibition | IMB Inflammasome Lab
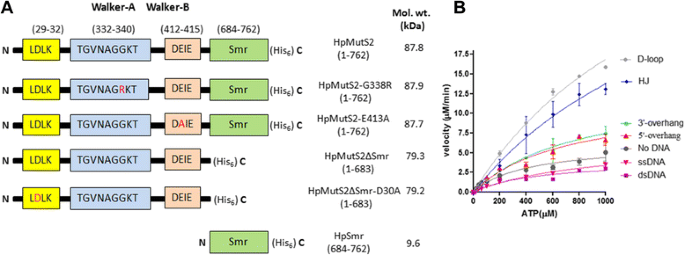
Mutations in the nucleotide binding and hydrolysis domains of Helicobacter pylori MutS2 lead to altered biochemical activities and inactivation of its in vivo function | BMC Microbiology | Full Text
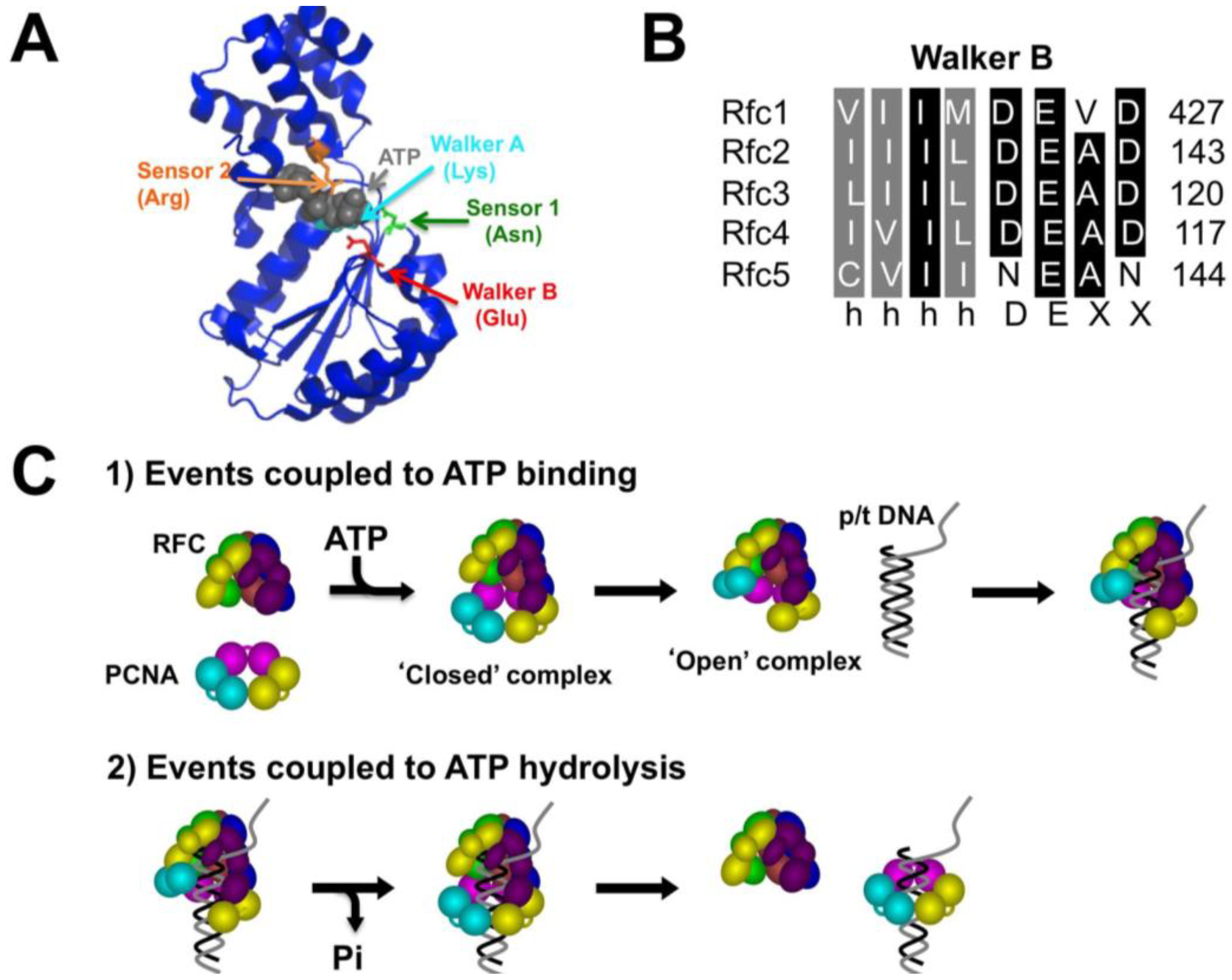
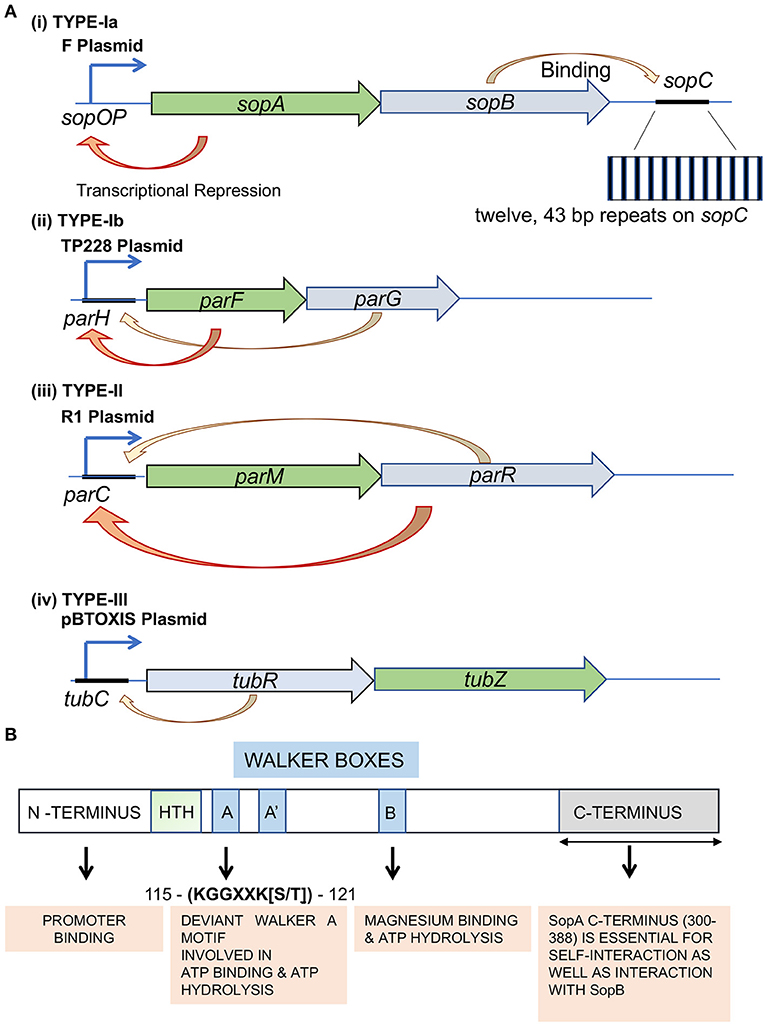
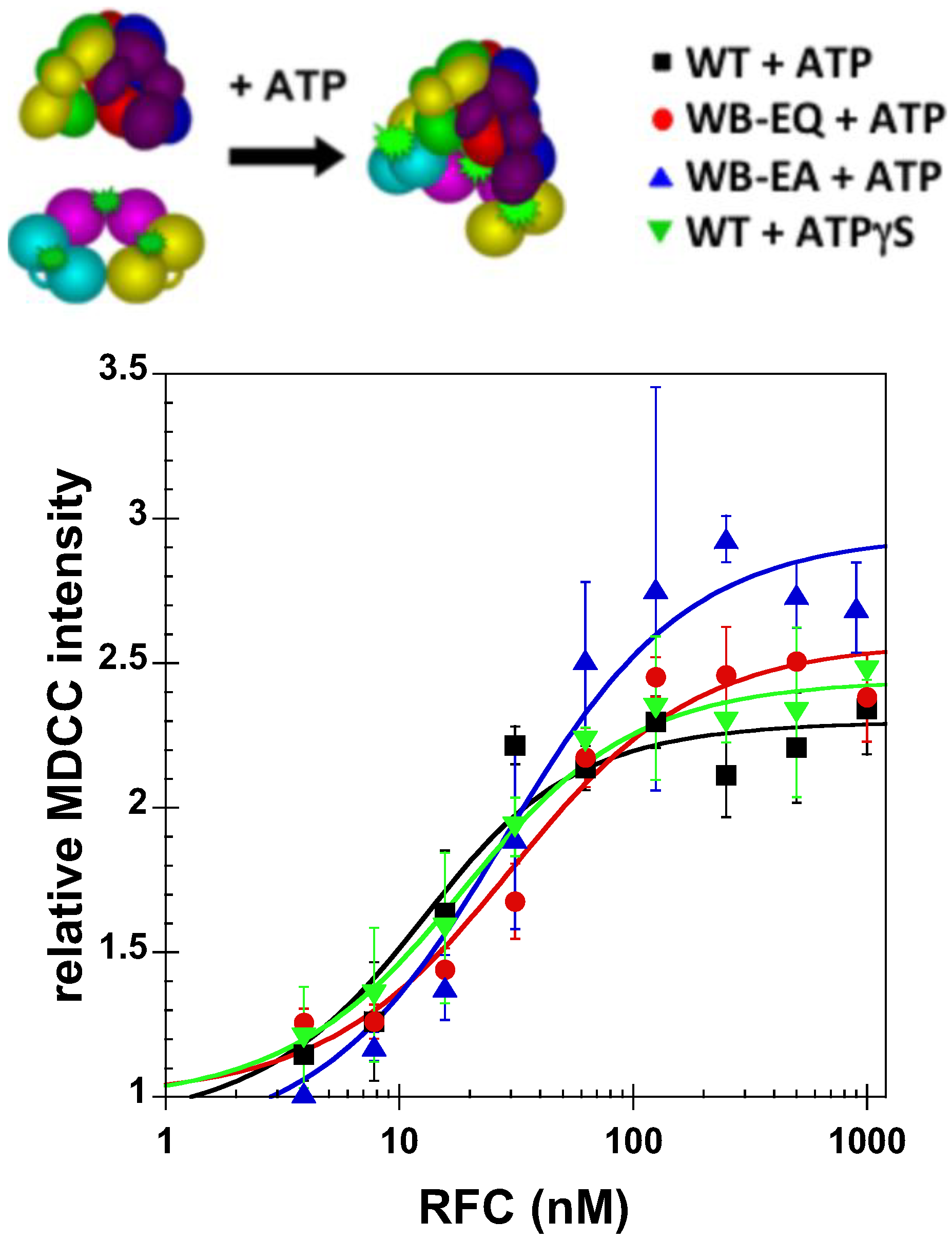
Genes | Free Full-Text | A Novel Function for the Conserved Glutamate Residue in the Walker B Motif of Replication Factor C
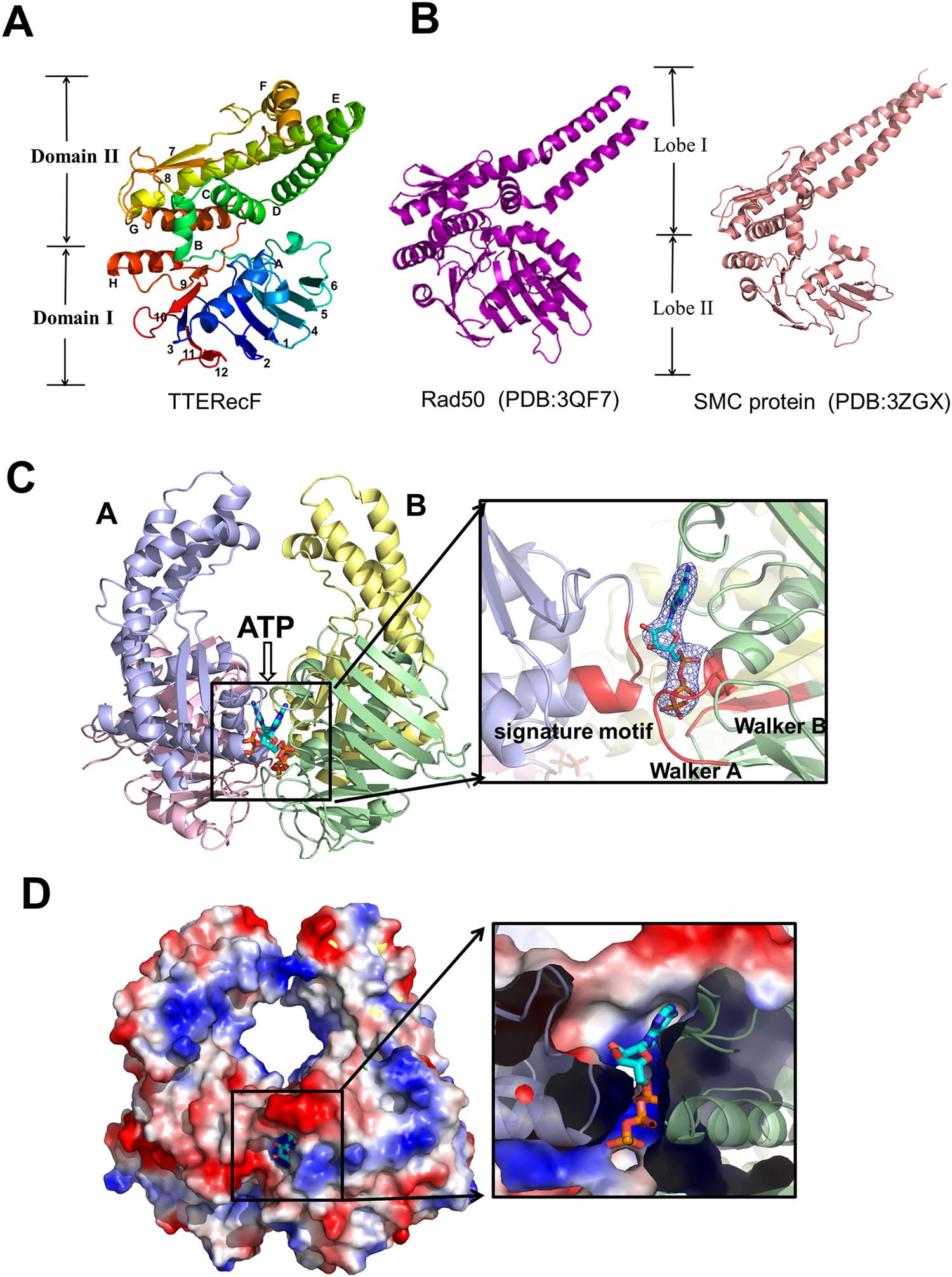
ATP-dependent conformational change in ABC-ATPase RecF serves as a switch in DNA repair | Scientific Reports

The putative Walker A and Walker B motifs of Rrp2 are required for the growth of Borrelia burgdorferi - Ouyang - 2017 - Molecular Microbiology - Wiley Online Library

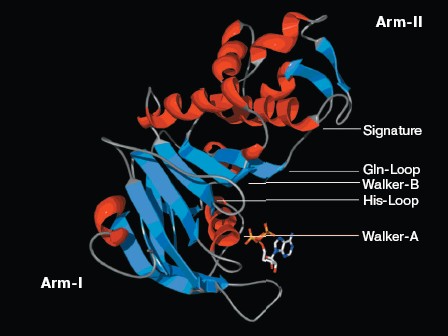
Visualization of NBD motifs: The NBD-NBD interface of a single NBD is shown in cartoon representation.

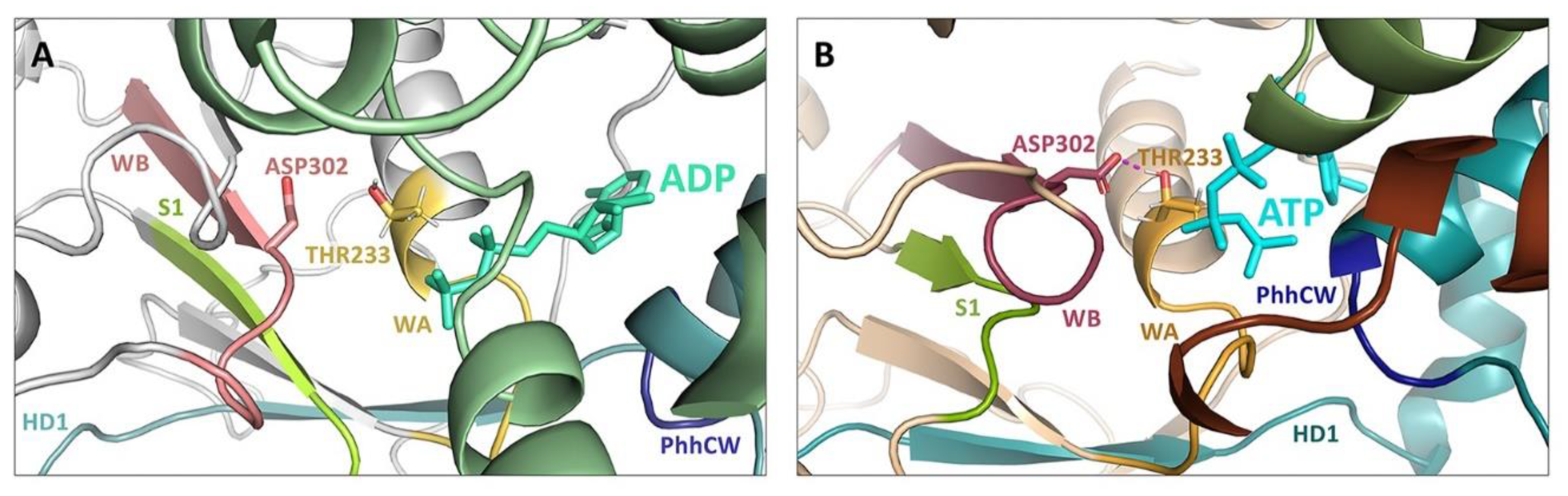
Model of the NOD2 nucleotide-binding site with an ADP molecule and conserved sequence motifs Walker A, Walker B, Sensor 1, GxP, and WH-His shown in sticks.
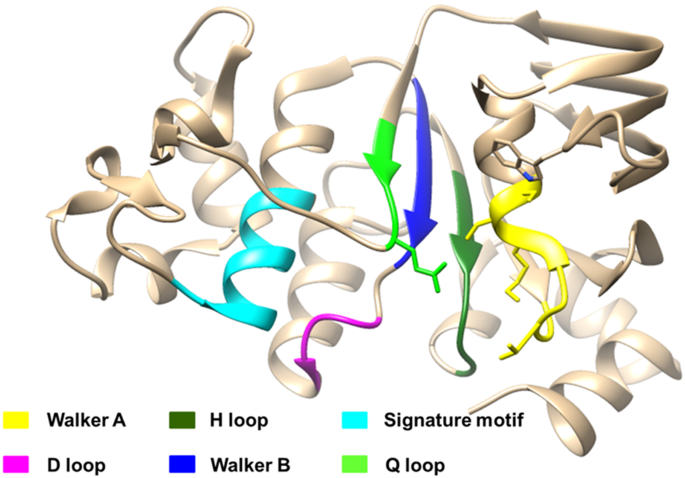
Targeting Nucleotide Binding Domain of Multidrug Resistance-associated Protein-1 (MRP1) for the Reversal of Multi Drug Resistance in Cancer | Scientific Reports
![SOLVED: The Walker A motif is protein structure element frequently observed in nucleotide hydrolases; which are enzymes that couple the energy released by breaking phosphate bonds with other reactions In the CH] SOLVED: The Walker A motif is protein structure element frequently observed in nucleotide hydrolases; which are enzymes that couple the energy released by breaking phosphate bonds with other reactions In the CH]](https://cdn.numerade.com/ask_images/9ecd6341cb4e4ea591d6564f0329d87f.jpg)
SOLVED: The Walker A motif is protein structure element frequently observed in nucleotide hydrolases; which are enzymes that couple the energy released by breaking phosphate bonds with other reactions In the CH]